What is gene ontology?
Gene ontology (GO) is an effort of the scientific community to organize the relationships of genes from various species and databases in terms of their biological processes, cellular component, and molecular function [1].
Organizing genes and genes products in this manner helps to simplify their understanding and communication. Sites such as AmiGO or QuickGO can be used to access the gene ontology of specific genes.
- Biological process ontology refers to some type of molecular event or process the gene/gene product is involved in
- Molecular function ontology refers to the specific action of a gene/gene product
- Cellular component ontology refers describes where in the cell a gene/gene product has activity [2]
Organizing genes and genes products in this manner helps to simplify their understanding and communication. Sites such as AmiGO or QuickGO can be used to access the gene ontology of specific genes.
CTLA4 gene ontology
Biological ProcessB Cell receptor signalling pathway
Immune response Negative regulation of B cell proliferation Negative regulation of regulatory T cell differentiation Positive regulation of apoptotic process Response to DNA damage stimulus |
Cellular ComponentClathrin-coated endocytic vesicle
External side of plasma membrane Golgi apparatus Integral to plasma membrane Perinuclear region of cytoplasm Plasma membrane |
Molecular FunctionProtein binding
Receptor activity |
CTLA4 gene ontology flowcharts
The following flowcharts obtained from QuickGO demonstrate the interactions of CTLA4 as defined by the gene ontology category.
|
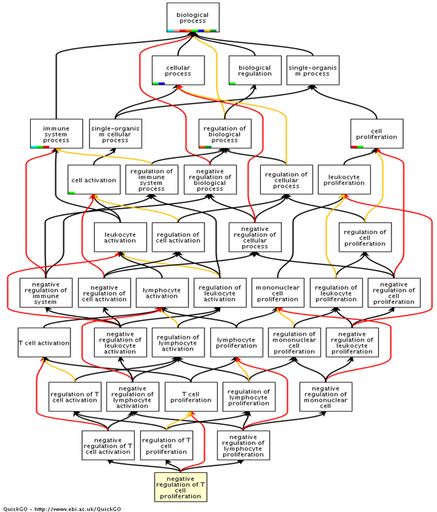
The biological process flowchart in figure 1 (below) shows the negative regulation of T cell proliferation by CTLA4. The complexity of the process is clear, with multiple layers of regulation and redundancy involving many different components of the immune response. It is clear how disruption of CTLA4 function can lead to an aberrant immune response.
|
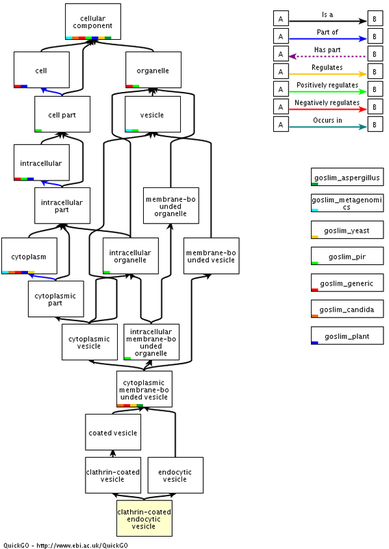
The cellular component flowchart in figure 2 (below) demonstrates the relationship of CTLA4 with clatherin coated endocytic vesicles. CTLA4 is primarily an intracellular protein, with tightly regulated surface expression. The interaction with clatherin coated endocytic vesicles is critical for intracellular and surface trafficking. Disruptions to this cellular component reduces the surface expression of CTLA4 resulting in disease status. [3]
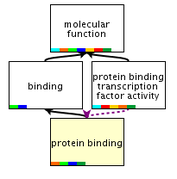
Protein binding (figure 3) is the primary molecular function of CTLA4 as the protein has no other intrinsic catalytic activity. The extracellular domain of CTLA4 binds costimulatory molecules to negatively modulate T cell activation. [3]
|
Analysis
The gene ontology of CTLA4 highlights its function as a membrane associated receptor responsible for the regulation of the immune response. Understanding how mutations in CTLA4 alter the cellular and molecular ontology will provide insight into the role of CTLA4 in disease phenotypes.
References
[1] The Gene Ontology Consortium. (2008). The gene ontology project in 2008. Nucleic Acids Research, 36(1):D440-4. doi: 10.1093/nar/gkm883
[2] GO: Quick tour - http://www.ebi.ac.uk/training/online/course/go-quick-tour/what-go
[3] Teft, W. A., Kirchhof, M. G., and Madrenas, J. (2006) A molecular perspective of CTLA-4 function. Annu. Rev. Immunol. 24, 65–97. doi: 10.1146/annurev.immunol.24.021605.090535
[2] GO: Quick tour - http://www.ebi.ac.uk/training/online/course/go-quick-tour/what-go
[3] Teft, W. A., Kirchhof, M. G., and Madrenas, J. (2006) A molecular perspective of CTLA-4 function. Annu. Rev. Immunol. 24, 65–97. doi: 10.1146/annurev.immunol.24.021605.090535