This web page was produced as an assignment for Genetics 677, an undergraduate course at UW-Madison.
Discussion
Grave's Disease is an autoimmune disorder characterized by the production of antibodies that target receptors of the thyroid gland. Among other genetic components, the CTLA4 gene has been linked to Grave's Disease susceptibility. CTLA4 is a key inhibitory receptor acting to modulate the immune activation of T cells. The protein consists of four exons encoding a leader peptide, extracellular domain, transmembrane region, and a cytoplamsic domain. [1] The protein structure is well conserved throughout mammalian species. The cytoplasmic domain is 100% conserved through different species, suggesting its structure is key to proper protein function. Disruption to any component of the protein could have physiologic effect, but only certain polymorphisms are observed. [2] Several studies have shown significant correlation between the 49A/G mutaion and Grave's disease. [1] Interestingly, the 49A/G mutation occurs in the signal peptide region of the peptide which is post translationaly cleaved. Observation of 49A/G mutant cells have showed altered binding to AP1 (clatherin coated adaptor protein 1) involved in the intracellular trafficking of CTLA4. However, the mechanisms by which the 49A/G mutation results in this phenotype is poorly characterized. [3] Further examination of intracellular trafficking will lead to a better understanding of CTLA4's role in Grave's Disease.
Experimental Approach
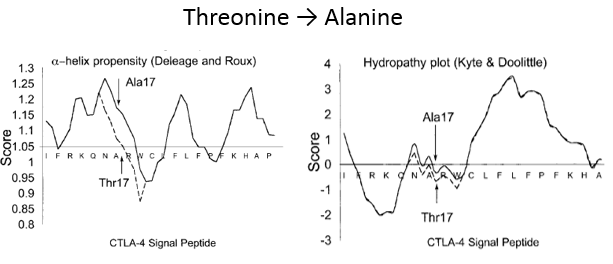
Understanding the 49A/G mutationThe 49A/G mutation causes an amino acid substitution at position 19 from threonine to alanine. This substitution slightly changes the hydropathy and alpha-helix propensity of the leader sequence. The peptide sequence is cleaved during translation of the mature peptide, so this mutation must cause some type of altered processing such as:
|
So how are the possible changes altering the protein trafficking of CTLA4 to result in a Grave's Disease phenotype?
|
Hypothesis: The 49A/G mutation impairs surface expression of CTLA4 but interrupting cellular trafficking.
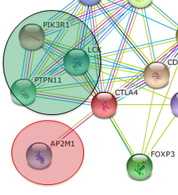
Experiment 1: Bioinformatics search for cellular trafficking proteinsSearching through protein interaction networks for human and its homologs I identified several proteins of interest. AP2 is a protein involved in the intracellular trafficking of CTLA4 and surface expression. There are also several protein modifiers (kinases and phosphorylases) that act on the cytoplasmic domain. Their interaction with CTLA4 influences stabilization of CTLA4 at the cell surface and influences the interaction with cellular trafficking components. [4]
|
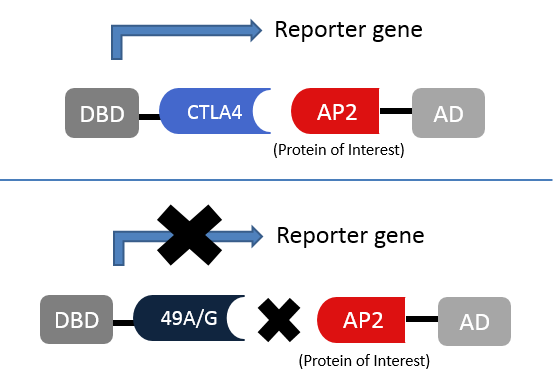
Experiment 2: Protein interactions altered by 49A/GA yeast two hybrid experiment can be performed to determine which protein interactions are disrupted by the 49A/G mutation. Even though many of the protein interactions are occurring at the membrane or in the cytoplasm, there are several different yeast two hybrid methodologies such as the split protein sensor system, split ubiquitin-based system or G-protein fusion yeast two hybrid. Coimmunoprecepitation should be coupled to the approach to reduce the rate of false positive interaction. This experimental can also be applied to identify new proteins the 49A/G mutation is interacting with.[5] Knowing which interaction are altered will help identify the mutations mechanism of action.
|
Experiment 3: MicroarrayIn order to completely understand the mechanism of the 49A/G mutation it would be helpful to understand the regulation of the proteins interacting with CTLA4. A microarray experiment may indicate the trafficking and modifying protein are expressed in conjuction with CTLA4 during immune stimulation, but this symmetry is disrupted in the 49A/G mutation.
|
Experiment 4: Identify post translational modifications
MALDI-TOF technology can be used to determine if the 49A/G mutation alters the glycosylation or phosphorylation of CTLA4 domains. Glycosylation events in the extracellular domain could alter dimerization, and phosphorylation events in the cytoplasmic domain could alter interaction with trafficking or modifying proteins. [6]
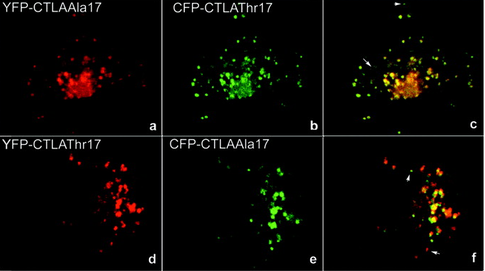
Experiment 5: Chemical genetic screen to restore protein functionI would perform a chemical genetics screen to identify a small molecule capable of restoring CTLA4 surface expression in a mutant cell line. The small molecule may directly cause increased CTLA4 surface expression or indirectly through interactions with trafficking/modifying proteins. [7] The surface expression of CTLA4 would be monitored as shown in figure 5, using florescently labeled protein and confocal microscopy.
When applied to the mouse model, the expected phenotype would be mouse survival with no lymphoproliferative disorders. |
Future Directions
This study will improve knowledge of CTLA4 interactions with trafficking components, which is critical to its function as an immune modulator. Future studies should examine the role of endosomal and lysosomal trafficking with respect to CTLA4. Mutations to CTLA4 may cause it to be directed away from the cellular membrane and into these intracellular compartments where it is sequester and eventually broken down.
If a small molecule is found to be effective in restoring CTLA4 surface expression, it could be applied to restoring the Grave's disease phenotype. The experimental findings could also be applied to other autoimmune disorders stemming from CTLA4 mutations or insufficient negative regulation of the immune response.
If a small molecule is found to be effective in restoring CTLA4 surface expression, it could be applied to restoring the Grave's disease phenotype. The experimental findings could also be applied to other autoimmune disorders stemming from CTLA4 mutations or insufficient negative regulation of the immune response.
|
Download my final presentation and experiments: |
|
|||||||||
References:
[1] Chistiakov D., Turakulov R., (2003). CTLA-4 and its role in autoimmune thyroid disease. J Mol Endocrinol, 31, doi: 10.1677/jme.0.0310021
[2] Teft, W. A., Kirchhof, M. G., and Madrenas, J. (2006) A molecular perspective of CTLA-4 function. Annu. Rev. Immunol. 24, 65–97. doi: 10.1146/annurev.immunol.24.021605.090535
[3] Anjos S, Nguyen A, Ounissi-Benkalha H,Tessier MC, Polychronakos C. 2002. A common autoimmunity predisposing signal peptide variant of the cytotoxicT-lymphocyte antigen 4 results in inefficient glycosylation of the susceptibility allele. J. Biol. Chem. 277:46478–86. doi: 10.1074/jbc.M206894200
[4] Valk E. Rudd C. Schneider H. 2008. CTLA-4 trafficking and surface expression. Trends in Immunology, 29, 272-279. doi:10.1016/j.it.2008.02.011
[5] Bruckener A. Polge C. Lentze N. Auerbach D. Schlattner U. 2009 Yeast Two-Hybrid, a Powerful Tool for Systems Biology. Int. J. Mol. Sci, 10, 2763-2788; doi:10.3390/ijms10062763
[6] April K.S. Salama and F. Stephen Hodi. 2011. Cytotoxic T-Lymphocyte Antigen-4. Clin Cancer Res; 17:4622-4628. doi: 10.1158/1078-0432.CCR-10-2232
[7] Lindsten, T., K. P. Lee, E. S. Harris, B. Petryniak, N. Craighead, P. J. Reynolds, D. B. Lombard, G. J. Freeman, L. M. Nadler, G. S. Gray, et al. 1993. Characterization of CTLA-4 structure and expression on human T cells. J. Immunol. 151:3489.
[2] Teft, W. A., Kirchhof, M. G., and Madrenas, J. (2006) A molecular perspective of CTLA-4 function. Annu. Rev. Immunol. 24, 65–97. doi: 10.1146/annurev.immunol.24.021605.090535
[3] Anjos S, Nguyen A, Ounissi-Benkalha H,Tessier MC, Polychronakos C. 2002. A common autoimmunity predisposing signal peptide variant of the cytotoxicT-lymphocyte antigen 4 results in inefficient glycosylation of the susceptibility allele. J. Biol. Chem. 277:46478–86. doi: 10.1074/jbc.M206894200
[4] Valk E. Rudd C. Schneider H. 2008. CTLA-4 trafficking and surface expression. Trends in Immunology, 29, 272-279. doi:10.1016/j.it.2008.02.011
[5] Bruckener A. Polge C. Lentze N. Auerbach D. Schlattner U. 2009 Yeast Two-Hybrid, a Powerful Tool for Systems Biology. Int. J. Mol. Sci, 10, 2763-2788; doi:10.3390/ijms10062763
[6] April K.S. Salama and F. Stephen Hodi. 2011. Cytotoxic T-Lymphocyte Antigen-4. Clin Cancer Res; 17:4622-4628. doi: 10.1158/1078-0432.CCR-10-2232
[7] Lindsten, T., K. P. Lee, E. S. Harris, B. Petryniak, N. Craighead, P. J. Reynolds, D. B. Lombard, G. J. Freeman, L. M. Nadler, G. S. Gray, et al. 1993. Characterization of CTLA-4 structure and expression on human T cells. J. Immunol. 151:3489.