This web page was produced as an assignment for Genetics 677, an undergraduate course at UW-Madison.
What are protein domains?
Protein domains are the structural units that can fold, function, and exist independently of the entire protein chain. Domains are often the active portion of a protein, defining the types of functions or interactions a protein will have. [1] For this reason, domains are often conserved through various species and through proteins with similar action. Domain analysis provides insight into a proteins mechanism of action. Several protein domain databases such as SMART, Prosite, and Pfam can be used to identify conserved domains and functional sites within a protein of interest.
Protein domains of CTLA4
|
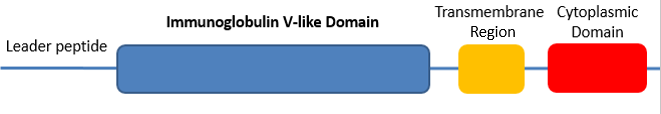
There are two known protein domains of CTLA4: the immunoglobulin domain, and the cytoplsamic domain. The protein databases were only able to confidently identify the immunoglobulin domain, with a slight discrepancy in the domain length. [2]
Immunoglobulin Domain: Ig-like domains resemble the antibody variable domain and are found in a diverse number of proteins. [3] In CTLA4, the domain is responsible for binding costimulatory ligands in the extracellular space. The domain is well conserved across homologs. Mutations to this domain may alter ligand binding leading to disease status. [2] Cytoplasmic Domain: The domain was not identified using the protein databases, but has been experimentally identified. The cytoplasmic domain has no intrinsic catalytic activity, but it interacts with a number of kinases and phosphorylases to initiate downstream effects. This domain is 100% percent conserved through mammalian species, indicating the proper function of this domain is critical to overall protein function. [2] Leader Peptide: The first 38 amino acids comprise the leader peptide that interacts with signal recognition particles to direct the mRNA sequence to the endoplasmic reticulum for translation. The signal peptide is cleaved during translation and is therefore not part of the mature protein. However, a mutation in the peptide sequence at posistion 17 from threonine to alanine has been significantly linked to Grave's disease. [2] Transmembrane Region: This region (often referred to as a domain) transverses the lipid membrane of cells. It consists of a helical structure. [2] |
What is a protein motif?
A motif is a short sequence of nucleotides or amino acids that can be distinguished as having some type of biological activity. The biological function of the motif may also imply conservation through different species. [4] Programs such as MOTIF can identify protein sequence motifs through multiple sequence alignments. MEME is an additional tool used to created sequence logos that help determine motif conservation.
Protein motifs of CTLA4
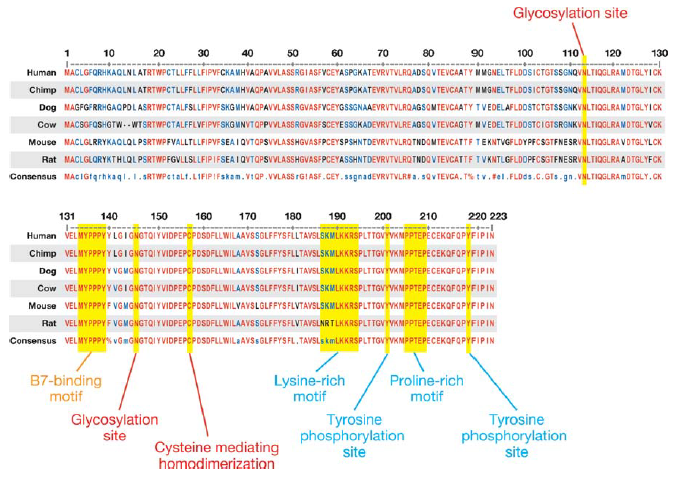
MOTIF was unable to identify any specific motif, but did identify the Ig-like domain. The figure below was obtained from a literature search and represents the alignment of homologous CTLA4 protein sequences. Red amino acids are conserved; blue amino acids have similar biochemical properties. Areas in yellow are possible motifs due to their potential functional relevance. The noted glycosylation sites are important for the dimerization of CTLA4. The B7 binding motif in the Ig-like domain is a point of contact between CTLA4 and its B7-family ligands. There are several motifs in the extremely well conserved cytoplasmic domain. These motifs play are role in the stabilization of CTLA4 at the cell membrane and have roles in downstream signalling. [2]
References
[1] Introduction to protein classification as the EBI: http://www.ebi.ac.uk/training/online/course/introduction-protein-classification-ebi/protein-classification/what-are-protein-domains
[2] Teft, W. A., Kirchhof, M. G., and Madrenas, J. (2006) A molecular perspective of CTLA-4 function. Annu. Rev. Immunol. 24, 65–97. doi: 10.1146/annurev.immunol.24.021605.090535
[3] Interpro: http://www.ebi.ac.uk/interpro/entry/IPR013106;jsessionid=ACFE35EFB50078459D0DEABD9C7B3ABA
[4] Bork, P., Koonin, E.V. (1996). Protein sequence motifs. Structural Biology, 6(3):366-76. doi: 10.1016?S0959-440X(96)80057-1
[2] Teft, W. A., Kirchhof, M. G., and Madrenas, J. (2006) A molecular perspective of CTLA-4 function. Annu. Rev. Immunol. 24, 65–97. doi: 10.1146/annurev.immunol.24.021605.090535
[3] Interpro: http://www.ebi.ac.uk/interpro/entry/IPR013106;jsessionid=ACFE35EFB50078459D0DEABD9C7B3ABA
[4] Bork, P., Koonin, E.V. (1996). Protein sequence motifs. Structural Biology, 6(3):366-76. doi: 10.1016?S0959-440X(96)80057-1