This web page was produced as an assignment for Genetics 677, an undergraduate course at UW-Madison.
Identifying Protein Interactions
A protein-protein interaction occurs when one protein binds another in order to carry out some biological process. These types of interactions can be hard to characterize due to the highly dynamic nature of proteins and the complexity of some interactions. A variety of methods such as yeast-two hybrid, TAP-tag, coimmunoprecipitation, and affinity chromatography (to name a few) can be used to identify different protein interactions. [1] Computer models, such as String, compile all know protein interactions from various sources into protein interaction maps. These maps give insight into the cellular role and regulation of a protein of interest, even leading to new understanding of protein relationships.
CTLA4 Protein Interactions
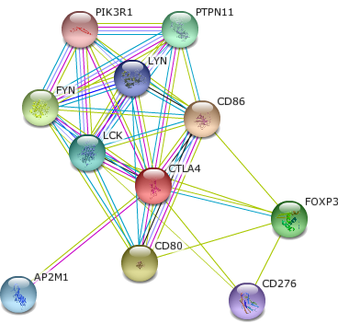
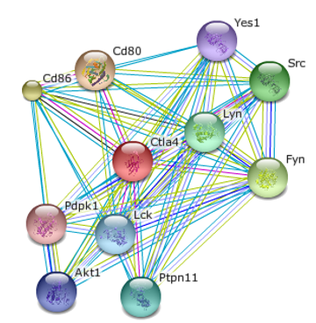
String was used to develop the interaction networks for human and mouse CTLA4. The validation method of each interaction is indicated by the color of the connecting line.
Analysis
There are several types of protein that CTLA4 interacts with that clarify its role as an immune modulator. CD86, CD80, and CD276, are involved in the costimulatory signal for T-cell activation, serving as ligands for CTLA4. There are also a series of phosphatases and kinases that likely interact with the cytoplasmic tail of CTLA4 to produce downstream effects. Several proteins act in the regulation of transcription, indicating a cell can control the expression of CTLA4 during immune stimulation. There is one protein of particular interest, AP2M1, a sub unit of the adaptor -related protein complex 2 that is responsible for intracellular trafficking of CTLA4. [2] It is known the surface expression of CTLA4 is tightly controlled, therefore any disruption in cellular trafficking may result in disease status. [3] Developing a deeper understanding of the components of CTLA4 intracellular and membrane trafficking may lead to an understanding of autoimmune disorders stemming from defective immune regulation. See [Future Directions] to see how CTLA4 mutation my lead to defective membrane trafficking and Grave's Disease.
The mouse homolog of CTLA4 is very similar to human CTLA4, with well conserved extracellular and cytoplasmic domains. It comes as no surprise that the protein interaction networks are also very similar, sharing similar protein interactions with ligands, phosphorylases/kinases, and transcriptional factors. There is no mouse AP2 trafficking protein shown, but experimental papers point to similar trafficking proteins in mice. [4] Many of the mouse interactions are not experimentally verified, indicating the role of CTLA4 has not been completely explored in mice. Due to the similarity in protein sequence and interaction network, the mouse still serves as a strong model organism.
The mouse homolog of CTLA4 is very similar to human CTLA4, with well conserved extracellular and cytoplasmic domains. It comes as no surprise that the protein interaction networks are also very similar, sharing similar protein interactions with ligands, phosphorylases/kinases, and transcriptional factors. There is no mouse AP2 trafficking protein shown, but experimental papers point to similar trafficking proteins in mice. [4] Many of the mouse interactions are not experimentally verified, indicating the role of CTLA4 has not been completely explored in mice. Due to the similarity in protein sequence and interaction network, the mouse still serves as a strong model organism.
References
[1] Phizicky EM, and Fields S,. (1995) Protein-protein interactions: Methods for detection and analysis. Microbiology Review. 59, 94-123
[2] Shiratori, T., Miyatake, S., Ohno, H., Nakaseko, C., Isono, K., Bonifacino, J. S., and Saito, T. (1997) Tyrosine phosphorylation controls internalization of CTLA-4 by regulating its interaction with clathrin-associated adaptor complex AP-2. Immunity 6, 583–589
[3] Valk E. Rudd C. Schneider H. 2008. CTLA-4 trafficking and surface expression. Trends in Immunology, 29, 272-279. doi:10.1016/j.it.2008.02.011
[4] Ling V, Wu PW, Finnerty HF, Sharpe AH, Gray GS, Collins M. (1999). Complete sequence determination of the mouse and human CTLA4 gene loci: cross-species DNA sequence similarity beyond exon borders. Genomics. Sep 15;60(3):341-55.
[2] Shiratori, T., Miyatake, S., Ohno, H., Nakaseko, C., Isono, K., Bonifacino, J. S., and Saito, T. (1997) Tyrosine phosphorylation controls internalization of CTLA-4 by regulating its interaction with clathrin-associated adaptor complex AP-2. Immunity 6, 583–589
[3] Valk E. Rudd C. Schneider H. 2008. CTLA-4 trafficking and surface expression. Trends in Immunology, 29, 272-279. doi:10.1016/j.it.2008.02.011
[4] Ling V, Wu PW, Finnerty HF, Sharpe AH, Gray GS, Collins M. (1999). Complete sequence determination of the mouse and human CTLA4 gene loci: cross-species DNA sequence similarity beyond exon borders. Genomics. Sep 15;60(3):341-55.